Cetacea is the most diverse group of current living aquatic mammals and includes 93 species. Cetaceans breathe air, despite their multiple adaptations to life in water, and therefore represent a very interesting benchmark to test whether this array of adaptations has left any signatures on mitochondrial tRNAs. In present paper, the evolution of mitochondrial tRNAs was investigated in the Cetacea, a clade of Cetartiodactyla that returned to the water and consequently adapted its metabolism to an environment different from that of its mainland ancestors.
TRNAs play a key role in mitochondrial activity thus, it is plausible that they experienced strong evolutionary constraints, particularly concerning their structural integrity, possibly further reinforced by the fact that there is a single tRNA for most amino acids. In this latter case, however, they are the product of duplication/multiplication processes and do not represent distinct codon families. Occasionally, multiple copies of the same tRNA are present in animal mtDNAs. Leucine and Serine are the only known exceptions and possess two tRNAs belonging to two distinct families (i.e., trnL1 and tnL2, and trnS1 and trnS2, respectively).

For most amino acids, a single tRNA represents the whole codon family.
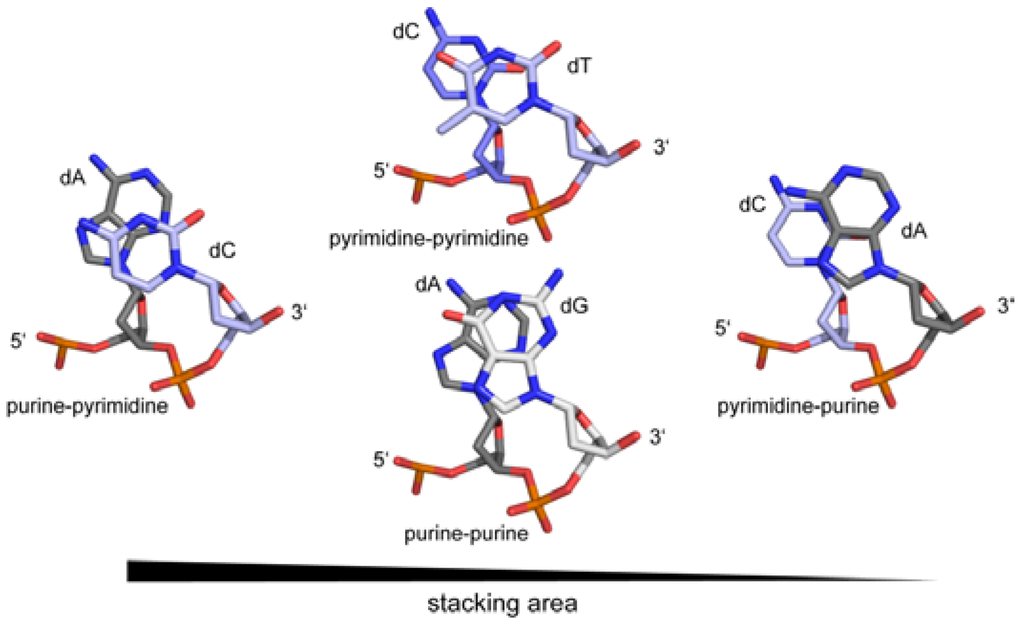
#IN DNA, A PURINE MUST ALWAYS PAIR WITH A PYRIMIDINE AND VICE VERSA IN ORDER TO ENSURE THAT CODE#
In most Metazoa, the mitochondrial genome (mtDNA) contains 22 tRNAs genes (hereafter named trnX, where X is the single letter IUPAC code for the corresponding amino acid). Mitochondrial tRNAs are fundamental components of the translational machinery of the mitochondrion, which is the powerhouse of the eukaryotic cell. This latter effect was particularly evident in toothed whales that either live in freshwater or are deep divers. Finally, it appears that the evolution of mitochondrial tRNAs was also shaped by the environments in which the Cetacean taxa differentiated.

Strong heterogeneity was observed among the different species of Cetacea. In particular, the range of variation and the fluctuation of these parameters affected the fate of single tRNAs. Our data suggested that the composition, AT-skew, and GC-skew of the tRNA stems were the main factors influencing the substitution process. Our analyses showed that the substitution pathways in the stems of different tRNAs were influenced by various factors, determining a molecular evolution that was unique to each of the 22 tRNAs. Our analysis focussed on identifying the factors that influenced the evolution of Cetacea tRNA double-helix elements, which play a pivotal role in the formation of the secondary and tertiary structures of each tRNA and consequently manipulate the whole translation machinery of the mitochondrion. In the present paper, the evolution of mitochondrial tRNAs was investigated in the Cetacea, a clade of Cetartiodactyla that retuned to water and thus had to adapt its metabolism to a different medium than that of its mainland ancestors. The mitochondrion is the power plant of the eukaryotic cell, and tRNAs are the fundamental components of its translational machinery.
